GMS:RT3D
| RT3D | |
|---|---|
| Model Info | |
| Model type | multi-species reactive transport |
| Developer | Battelle Pacific Northwest National Laboratory |
| Documentation | RT3D Manual v1 RT3D Manual v2.5 |
| Tutorials | RT3D Tutorials |
RT3D (Reactive multi-species Transport in 3-Dimensional groundwater systems) is a model developed by the Pacific Northwest National Laboratory. RT3D is a modified version of MT3DMS that utilizes alternate chemical reaction packages. Numerous pre-defined reactions are available and an option is provided for creating user-defined reactions. RT3D is well-suited for simulating natural attenuation and bioremediation.
Since RT3D is a modified version of MT3DMS, most of the input to RT3D is identical to the input required for MT3DMS. Thus, the RT3D interface is contained within the MT3DMS menu in the 3D Grid module. In the Basic Transport Package dialog, an option is provided for selecting the current model as either MT3DMS, RT3D, or SEAM3D. A number of options in the interface then change based on which model is selected.
Since much of the RT3D interface is identical to the MT3DMS Interface, only the portions of the interface which are unique to RT3D are described in this help file.
The RT3D model can be added to a paid edition of GMS.
Contents
Basic Transport Package
The first step in defining an RT3D simulation is to define the data required by the Basic Transport (BTN) package. The options in the Basic Transport Package dialog unique to RT3D are as follows:
Packages
The Packages button brings up the Packages dialog. If the RT3D model is the current model and the Chemical Reaction package option is selected, one of the RT3D reactions must be selected from the pull-down list. The first nine reaction modules are pre-defined (specified as the IREACT value):
- No reaction (tracer transport)
- Inst. Aerobic Deg. of BTEX
- Inst. Deg. of BTEX w/ MEA
- Kinetic-Limited Deg. of BTEX w/ MEA
- Rate-Limited Sorption Reactions
- Double Monod Model
- Sequential Decay Reactions
- Aerobic/Anaerobic PCE/TCE Dechlorination
- Test Model #1
- Test Model #2
If one of these reactions is selected, the names of the species and the names of the reaction parameters are automatically determined by GMS. The last reaction is a user-defined reaction. If this option is selected, a list of species and list of reaction parameters must be specified.
For more information on each reaction package, see the RT3D Reaction Packages article.
Define Species
For most of the reaction package options, once the reaction package is selected, the list of species used by the package is automatically initialized by GMS. However, if the user-defined reaction package is selected, the Define Species button is undimmed in the Basic Transport Package dialog and a list of species must be manually defined before any concentrations are assigned to sources/sinks.
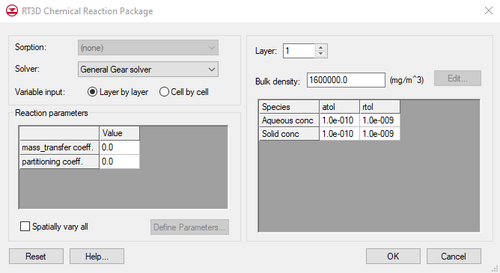
Chemical Reactions Package
The Chemical Reactions Package dialog utilized by RT3D is different from the MT3DMS Chemical Reactions dialog. The items in the dialog unique to RT3D are as follows:
Solver
If the selected reaction package is a kinetic reaction, several solvers are available for the solution of the chemical reaction equations. The desired solver should be selected from the Solver pull-down list. When one of these solvers is used, an absolute and relative tolerance must be specified for each of the mobile species using the atol and rtol parameters.
Reaction Parameters
With each reaction package, a set of reaction parameters must be defined. The method used to edit the reaction parameters depends on whether the reaction is a pre-defined reaction or a user-defined reaction.
Pre-Defined Reactions
For the pre-defined reactions, the number of reaction parameters and the names of the parameters are fixed. If the Spatially vary all toggle is off, a single value is entered for each parameter using the edit field below the parameter list. If the Spatially vary all toggle is on, an array of values is entered for each parameter using the Edit button. In this case, the cell-by-cell parameter values can also be edited using the Cell Attributes command.
User-Defined Reactions
For user-defined reactions, the list of reaction parameters must be defined using the Define Parameters button. This button brings up the Define Parameters dialog. This dialog functions similarly to the Define Species dialog. The New and Delete buttons are used to add and remove items from the list. The Import button is used to read a previously defined list of reaction parameters from an RT3D super file (*.rts).
With the pre-defined reactions, the reaction parameters are either all spatially variable or all constant. However, with the user-defined reaction option, selected parameters may be designated as spatially variable while others are designated as constant. The variable/constant status of a parameter is selected using the Spatially variable toggle in the Define Parameters dialog.
RT3D Files
Below is a table of available RT3D files.
- For more information on these files see pages 11–38 of the original RT3D online documentation ([1]).
| Name | Description |
|---|---|
| ADV | Advection Package File |
| BTN | Basic Transport File |
| CON | Concentration Output File |
| DSP | Dispersion Package File |
| DSS | Data Super File |
| MTR | GMS Results File |
| OUT | Output File |
| RCT | Chemical Reaction Package File |
| RTS | RT3D Super File |
| SSM | Sources/Sink Mixing File |
| GMS – Groundwater Modeling System | ||
|---|---|---|
| Modules: | 2D Grid • 2D Mesh • 2D Scatter Point • 3D Grid • 3D Mesh • 3D Scatter Point • Boreholes • GIS • Map • Solid • TINs • UGrids | |
| Models: | FEFLOW • FEMWATER • HydroGeoSphere • MODAEM • MODFLOW • MODPATH • mod-PATH3DU • MT3DMS • MT3D-USGS • PEST • PHT3D • RT3D • SEAM3D • SEAWAT • SEEP2D • T-PROGS • ZONEBUDGET | |
| Aquaveo | ||